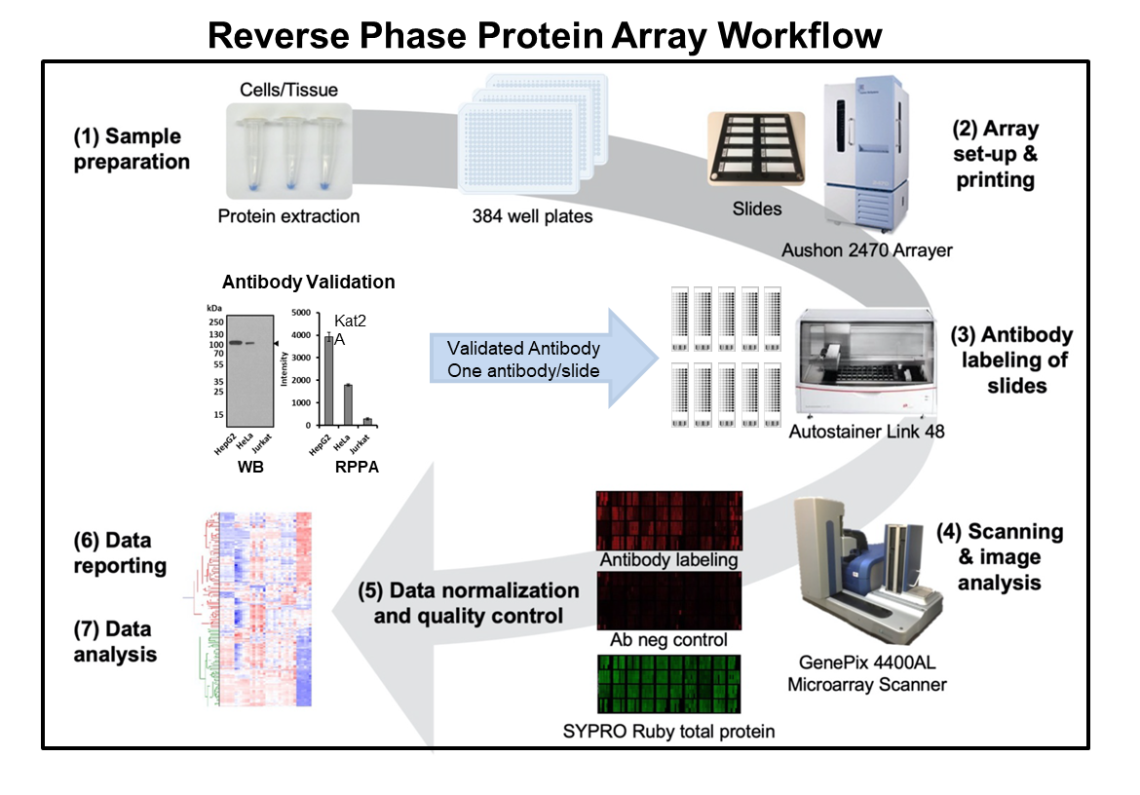
Flow chart illustration the Reverse Phase Protein Array process.
Coarfa C, Grimm SL, Rajapakshe K, Perera D, Lu HY, Wang X, Christensen KR, Mo Q, Edwards DP, Huang S. Reverse-Phase Protein Array: Technology, Application, Data Processing, and Integration. J Biomol Tech. 2021 Apr;32(1):15-29. doi: 10.7171/jbt.21-3202-001. PMID: 34025221; PMCID: PMC7861052.
Reverse Phase Protein Array
Reverse phase protein array (RPPA) is a high-throughput technology that performs protein assays on thousands of samples simultaneously, including tissue and cell lysates, serum, plasma or other body fluids. This protein array platform measures levels of protein expression, as well as protein modifications such as phosphorylation. Protein lysates are arrayed as microspots on nitrocellulized coated glass slides and probed with highly specific antibodies that have been validated for RPPA. Each microspot contains the whole proteome repertoire of the tissue or cell. This enables the determination of a set of parameters in large collections of tissue or cell samples.
We currently have an inventory of 264+ validated antibodies that cover multiple total proteins and phosphoproteins in the following protein pathways or functional protein groups: epithelial-mesenchymal transition (EMT), stem cells, apoptosis, DNA damage, Autophagy, proliferation and cell cycle, growth factor receptors, cytokines/STATs, nuclear receptors/transcriptional regulatory proteins, autophagy and metabolomics enzymes. We continually work together with BCM investigators to build RPPA assays for new protein pathways of interest and to validate the required antibodies. We are also working to be able to do analysis with various types of samples including cell/tissue lysates, macro-dissected tumor samples, laser capture micro-dissected (LCM) samples, small numbers of isolated stem cells or other rare cell types, and body fluids such as serum and urine. A novel application we have developed is to analyze isolated protein complexes. We are also developing epigenetics RPPA platform. We have developed in-house normalization and statistical analyses algorithms with the Multi-Omics Data Analysis.
iLabs
iLabs Service Request








